In the last couple of years, researchers have been encouraged to start using metadata when they collect data for clinical research. Funding parties, such as ZonMW, even require you to use and describe metadata in your data management plan. But what are metadata and why are they so important? In this article we will explain what metadata are and how you can apply them in your next research project.
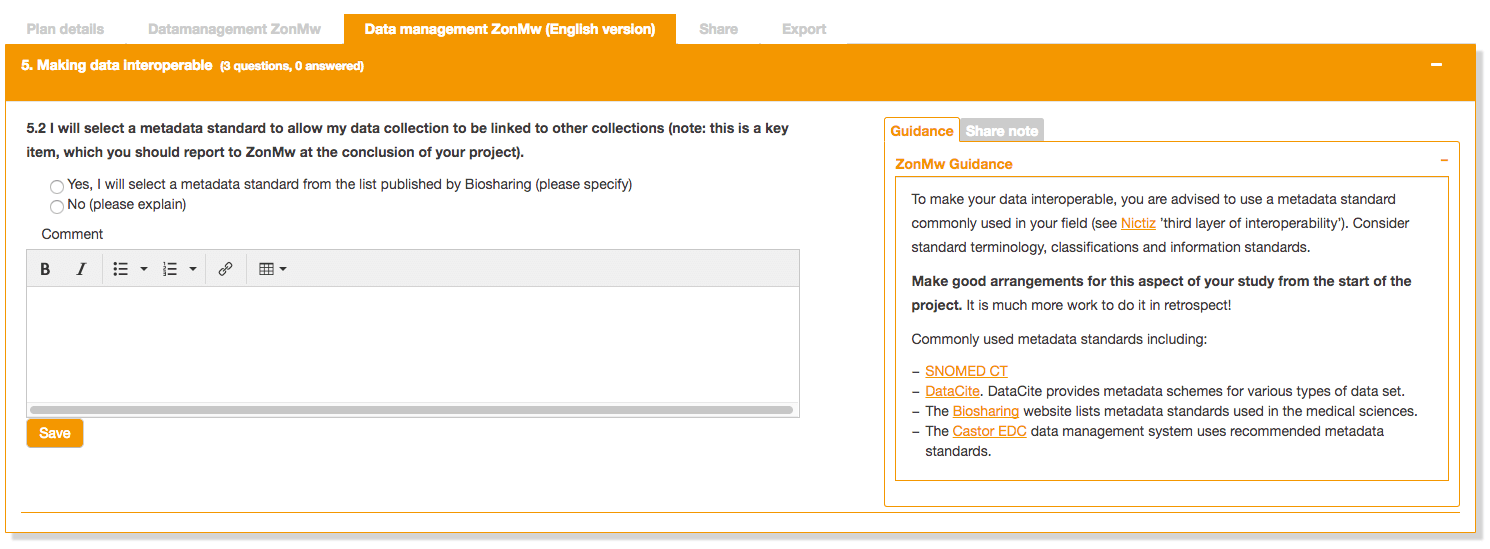
Metadata standards in the ZonMW Data Management Plan
What are metadata?
Metadata are data that provide information about other data, generally a dataset. These data, in combination with documentation, help you, your colleagues, and computers to comprehend the meaning behind the data in the datasets. Metadata also enables interoperability (i.e. the ability to exchange data and interpret that data) since they provide a clear definition of the data that can be used to interpret them. Most metadata in medical datasets are added using ontologies.
What are ontologies?
Ontologies contain formal specifications of terms and relationships between them. You can think of them as as dictionaries, where every word in the dictionary (a concept) has a unique identifier (concept ID) and is described by other words from that dictionary (definitions and relationships). Since all concepts in the ontology are unique and defined by their relations with other concepts, they give a good explanation of what the concept means and how it is related to other concepts.
Example
In SNOMED CT (an ontology) lying systolic blood pressure is expressed in concept 407556006 | Lying systolic blood pressure (observable entity) |, where 407556006 is the concept ID and Lying systolic blood pressure (observable entity) is the description.
Why should I use metadata?
The usage of metadata helps you, but also other researchers, to understand what the data in your dataset is about. For example, if you have not looked at your dataset for five years and are starting to work with it again, the metadata helps you to understand the data, instead of you having to figure it out all over again. Moreover, metadata makes linking variables between datasets easier and, therefore, helps you to reuse or share data with colleagues.
How should I use metadata?
You can link the unique identifiers of concepts from ontologies to the variable names and option list items that you use in your dataset. By ‘coding’ these variables and options, they become readable by computers and also by other researchers, since these codes uniquely define what the variables or options are about.
What ontology should I use?
This is dependent on your field of research, and may result in the use of a combination of ontologies. Ontologies such as SNOMED CT and LOINC contain concepts for a broad range of fields, but do not include all field-specific concepts like gene expressions that are included in the Gene Ontology or phenotypic abnormalities in the Human Phenotype Ontology. The NCBO BioPortal and FAIRsharing give an overview of all the ontologies that are in use in your field of research and also shows updates frequencies for each ontology.
Your hospital or research institute might also use institutional lab codes that you can employ to code your variables. However, instead of only using these codes, please consider coding them with international ontologies (e.g. LOINC for lab data) for ease of sharing or reusing data later on.
Example
During a study that looks at hypertension in The Netherlands, you measure the blood pressure of the patient while the patient is lying down. You include this measurement in your CRF with two variable names: ‘Systolische bloeddruk’ (Systolic blood pressure) and ‘Diastolische bloeddruk’ (Diastolic blood pressure). At this point, your variables do not indicate how the measurement is performed and, since they are in Dutch, foreign researchers might not understand the meaning of the variables.
To make sure that your data is interoperable, you can add metadata to these variables that explain their meaning. SNOMED CT contains concepts that express these variables well:
-
- 407556006 | Lying systolic blood pressure (observable entity) |
- 407557002 | Lying diastolic blood pressure (observable entity) |
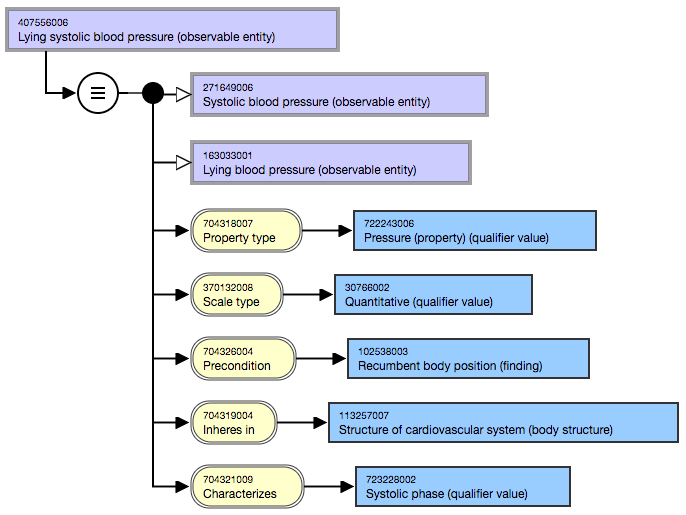
As you can see in the figure below, the Lying systolic blood pressure concept explains that this blood pressure is a systolic blood pressure and a lying blood pressure (purple boxes) and this concept has several attributes relationships (yellow) that relate to other concepts (blue).
Diagram of SNOMED CT concept 407556006 | Lying systolic blood pressure (observable entity) |
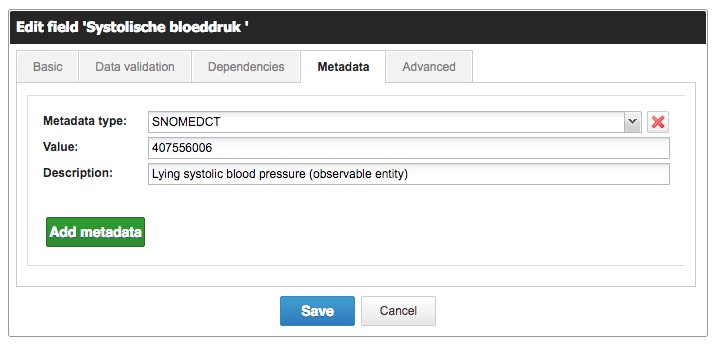
Looking back at your variables: you can add these SNOMED CT concepts as metadata on these variables.
Adding metadata to a field in Castor
If you export the structure of your study, you can see that the SNOMED CT concept is included in the metadata definition of this field. Then, if you want to share your dataset with other researchers, you can easily send this structure along and they will be able to interpret exactly what your variables are measuring. In the future Castor will also support other standardized formats to exchange your metadata.
<metadata>
<m_D1939C1D-5598-ABF8-7170-A031AD62C4D9>
<metadata_id>D1939C1D-5598-ABF8-7170-A031AD62C4D9</metadata_id>
<metadata_parent_id></metadata_parent_id>
<element_id>5ECBF6EB-24C1-1759-5F1E-43132FFFA40C</element_id>
<element_type>1</element_type>
<metadata_type>1</metadata_type>
<metadata_description>
Lying systolic blood pressure (observable entity)
</metadata_description>
<metadata_value>407556006</metadata_value>
</m_D1939C1D-5598-ABF8-7170-A031AD62C4D9>
</metadata>
...
<metadataTypes>
<mt_1>
<type_id>1</type_id>
<name>SNOMEDCT</name>
<description>SNOMED CT concepts</description>
</mt_1>
</metadataTypes>
Metadata definition of a field in the Castor structure export (XML)
Getting started yourself
Are you using Castor already and want to use metadata in your study? You can add a metadata type in the Settings tab and add metadata to your fields in the Form Builder. For more information on this, you can check our manual.
Sources:
1 http://www.sciencedirect.com/science/article/pii/S1042814383710083?via%3Dihub